Compound Discoverer 3.3 SP3 released
Compound Discoverer 3.3 SP3 software has just been released! We highly recommend upgrading to CD3.3SP3 as soon as possible. This new release contains bug fixes, database updates and some new features, but most importantly it also includes major performance improvements. To access your software from the download site, log in to your account here: https://thermo.flexnetoperations.com/
In the Release Notes you can find instructions for a “clean” install of CD3.3SP3. Please keep in mind that different versions of CD (e.g. 3.2 and 3.3) can coexist on the same PC. However this is not the case for service pack versions, for example 3.3.2 and 3.3.3. You need to uninstall your current CD3.3 before installing CD3.3 SP3.
Enhancements made to the Compound Discoverer 3.3 SP3 application:
- Performance improvements: Many computationally intensive data processing nodes, like Detect Compounds, Fill Gaps and Apply mzLogic™, are now executed 2 to 8 times faster, depending on the configuration of the data processing computer.
- Database updates:
- A new spectral library and mass list with 48,646 structures (Getzinger et. al. , Structure Database and In Silico Spectral Library for Comprehensive Suspect Screening of Per- and Polyfluoroalkyl Substances (PFASs). DOI: 10.1021/acs.analchem.0c04109.) are available for the identification of PFAS compounds. The updated PFAS workflow now also utilizes this database and spectral library.
- Human Metabolome Database (HMDB) metabolite databases (urine, serum, CSF, saliva, feces, sweat, all metabolites) are now available as mass lists under Lists & Libraries > Mass Lists.
- A new version of the mzCloud™ Offline Spectral Library (2023) is available under Lists & Libraries > Spectral Libraries.
- A new version of the Natural Products Atlas database (v2023_06) is available under Lists & Libraries > Mass Lists.
- Molecular Networking: Additional information about compounds and connections can be displayed.
- Support for data acquired on Orbitrap Astral instruments.
- […] and more. For a detailed list of changes please see the Release Notes.
Rawrr… !
I just found this really cool R-package today: Rawrr by Tobias Kockmann and Christian Panse. This package allows direct access to Thermo Fisher Scientific’s RAW files within the R environment, by wrapping the RawFileReader. Also, a big thank you to the authors for making the installation of the package really easy. If you don’t have the RawFileReader dll’s installed, it will automatically download them for you.
Join us at our annual Compound Discoverer User Meeting in Bremen – December 12-13, 2023
We are delighted to invite you to our upcoming Compound Discoverer Software Users Meeting in Bremen, Germany!
Join us for an enriching session designed to cater to both newcomers and seasoned users. Our goal is to enhance your understanding of best practices, whether you’re just starting or looking to take your skills to the next level. Advanced users will have the opportunity to engage with software experts, unlocking the full potential of their data.
Stay informed about the latest advancements and cutting-edge technologies while connecting with fellow users.
Discover firsthand how Thermo Scientific Compound Discoverer Software is being utilized across diverse applications. Our keynote speaker, Lee Ferguson, Duke University, USA will be presenting: Non-targeted analysis of water pollutants using the Orbitrap Astral Mass Spectrometer. Don’t let this opportunity pass you by – seize the chance to boost your software proficiency and expand your professional network!
The registration fee is 250 euros plus VAT and includes dinner on Day 1 at Canova. Upon registering, you will receive an invoice via email for your payment convenience.
Full agenda and registration: www.thermofisher.com/DiscovererSoftwareUserMeeting
Save the date! Compound Discoverer Users Meeting in Bremen
Our annual Compound Discoverer Users Meeting will happen at exactly the same days as last year, December 12 and 13th, 2023 at the Atlantic Hotel in Bremen, Germany. More details will follow shortly!
Compound Discoverer Software virtual training workshops
We are pleased to invite you to our Autumn 2023 version of virtual activities to support our Compound Discoverer (CD) software users.
The workshops are designed to bring beginners up-to-speed with best practices as well as allow advanced users to interact with software experts and enquire how to get the most out of their data. Additionally, two dedicated sessions specialized in Metabolite/drug metabolite Identification (Met ID) and Gas Chromatography Mass Spectrometry (GCMS) data analysis will follow.
The content is delivered through online workshops, 2-3 hours each, via MS-Teams. The workshops feature live exercises on pre-distributed data, discussion of CD features, parameters, and optimized settings. The sessions are hands-on and highly interactive, so get the best experience by having CD 3.3 SP2 up and running. Click HERE to download a free 60-day trial version of the latest Compound Discoverer software release.
The sessions allow ample time for the participants to get their CD-related questions answered in real time. The workshops will be delivered in English, but participants can use MS Team`s instant subtitles in their preferred language.
The workshops will be offered for two different time zones.
North, Central, and South America
General Compound Discoverer software training:
Tuesday 26th September 12.30-15.30 EDT (NY time)
Wednesday 27th September 12.30-15.30 EDT (NY time)
Thursday 28th September 12.30-15.30 EDT (NY time)
Specialized data processing:
Met ID: Wednesday 5th October 12.30-15.30 EDT (NY time)
GCMS: Thursday 17th October 12.30-15.30 EDT (NY time)
How to register:
Click HERE for registration (AMER region).
Europe, the Middle East, and Africa
General Compound Discoverer software training:
Tuesday 26th September 10.00-13.30 CEST (Berlin time)
Wednesday 27th September 10.00-13.30 CEST (Berlin time)
Thursday 28th September 10.00-13.30 CEST (Berlin time)
Specialized data processing:
Met ID: Wednesday 4th October 10.00-13.30 CEST (Berlin time)
GCMS: Thursday 18th October 10.00-13.30 CEST (Berlin time)
How to register:
Click HERE for registration (EMEA region).
We are looking forward to meeting you at these workshops,
Compound Discoverer software team
New scripting node
Stable Isotope Labeling. Sometimes people would like to see the relative peak area for each isotopologue reported in addition to the label incorporation rate. There is now a scripting node available for that. It exports these information into an Excel spreadsheet, for compounds that are checked or all compounds if none are checked.
Compound Discoverer Users Meeting at ASMS
Compound Discoverer Users Meeting on Saturday, June 3rd 2023, 9-12am at ASMS in Houston, Texas.
Compound Discoverer 3.3 SP2 released
We are happy to announce that Compound Discoverer 3.3 SP2 software is now available for download. To access your software from the download site, log in to your account here: https://thermo.flexnetoperations.com/
Enhancements in the Compound Discoverer 3.3 SP2 application
• The Spectral Libraries library includes one additional mass spectral library: the Epoxidized Soybean Oil Library
AN001586, which is a mass spectral library generated from compounds identified in epoxidized soybean oil.
• The Mass Lists library includes four new mass lists:
– Three new mass lists for the identification of PFAS compounds: PFAS_NIST, PFAS_NEG, and Chemical List PFASSTRUCT-2022-04-20
– The Food Contact Chemicals database mass list (FCCDB_2022)
• The Mass Lists library also includes two updated mass lists:
– The LipidMaps Structure Database mass list has been updated to the latest version 2023-01-11.
– The Extractables and Leachables HRAM Compound Database – 2023 Update has been expanded to 2,662
compounds.
• The Compound Classes library includes the following two new compound class files for the identification of
PFAS compounds:
– PFAS Fine structure fragment_lib
– PFAS General from FluoroMatch Suite
• A new processing workflow template for identifying per- and polyfluoroalkyl substances (PFAS) has been added
to the set of templates for LC studies.
• The Transformations list includes one new transformation reaction: PFAS Chain Shortening (for PFAS
compounds).
• The limit for the number of compounds that you can export to the molecular networks viewer has been
increased to 3,000.
• A new tutorial for the mapping of biological pathways using the BioCyc Genome Database Collection has been
added to the set of user guides and tutorials that you can access from the Help > Manuals menu
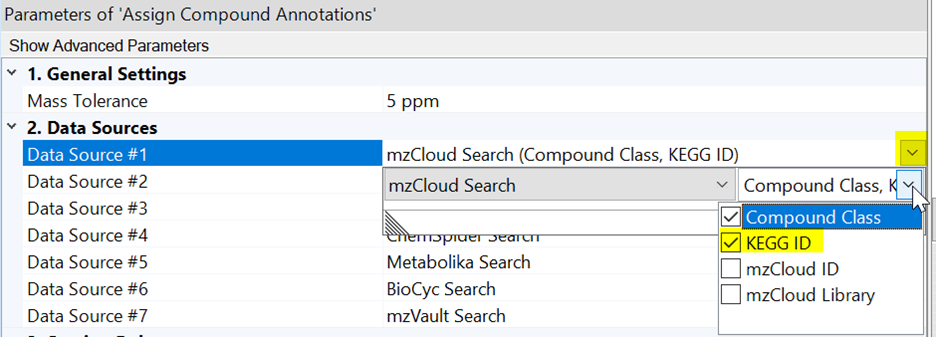
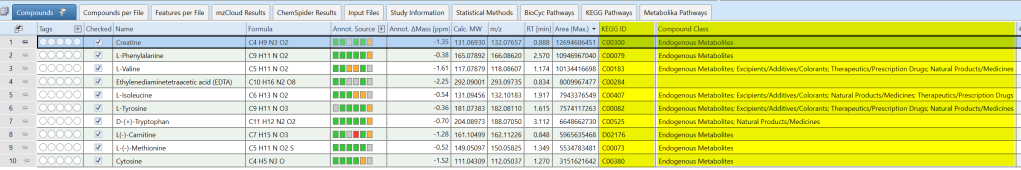
How to display KEGG IDs and Compound Class in the Compounds table
Compound Discoverer can display KEGG IDs or Compound Class as an additional column in the “Compounds” table for all compounds with hits in mzCloud™. To ensure desired IDs are displayed in your Compounds table, please edit the parameters of the “Assign Compound Annotation” node and select KEGG ID or Compound Class in the dropdown menu for the mzCloud Search.
BioCyc – Biological pathway mapping in Compound Discoverer
BioCyc has recently gained popularity as an excellent pathway mapping tool. The database accommodates metabolic networks for ~ 20,000 sequenced organisms across 3,105 pathways, and ~ 19,000 metabolites. Its superior informatics functions allow zoomable pathway diagrams featuring chemical structures and offer the key capability of overlaying user-obtained OMICS study results (e.g., fold change, peak area, or stable isotope labeling data) onto organism-specific metabolic pathways, producing data-oriented pathway representation and leading the way to successful biological interpretations.
Many academic organizations have institutional subscriptions https://biocyc.org/subscriber-list.shtml and BioCyc offers a free trial period for all Thermo Fisher Scientific users as well as a 20% discount for a one-year subscription. For more information, please visit https://biocyc.org/thermofisher.shtml
Attached is a tutorial showing you how to work with the BioCyc Pathways mapping feature in Compound Discoverer 3.3 service pack 2 (SP2) application.